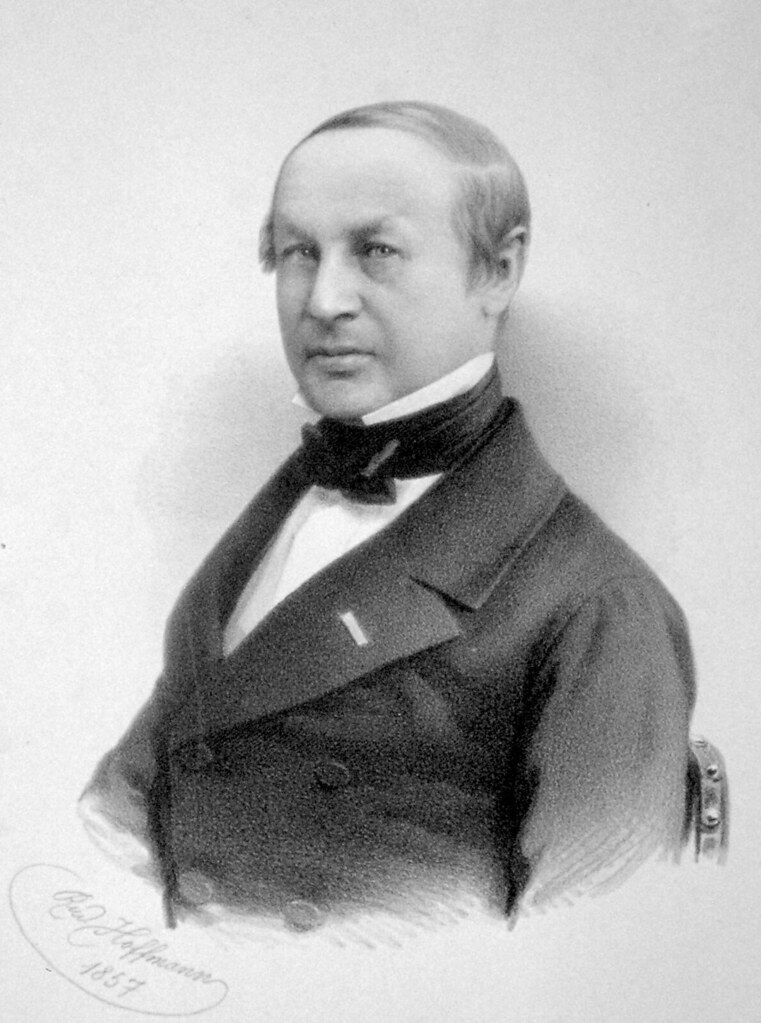![]()
Today is the 57th birthday of Chris White. Chris founded the yeast company White Labs in 1995 and he’s also on the faculty of the Siebel Institute. He’s also a fixture at virtually every brewing industry and homebrewing conference, and was kind enough to talk to my SSU beer appreciation class about yeast. Join me in wishing Chris a very happy birthday.
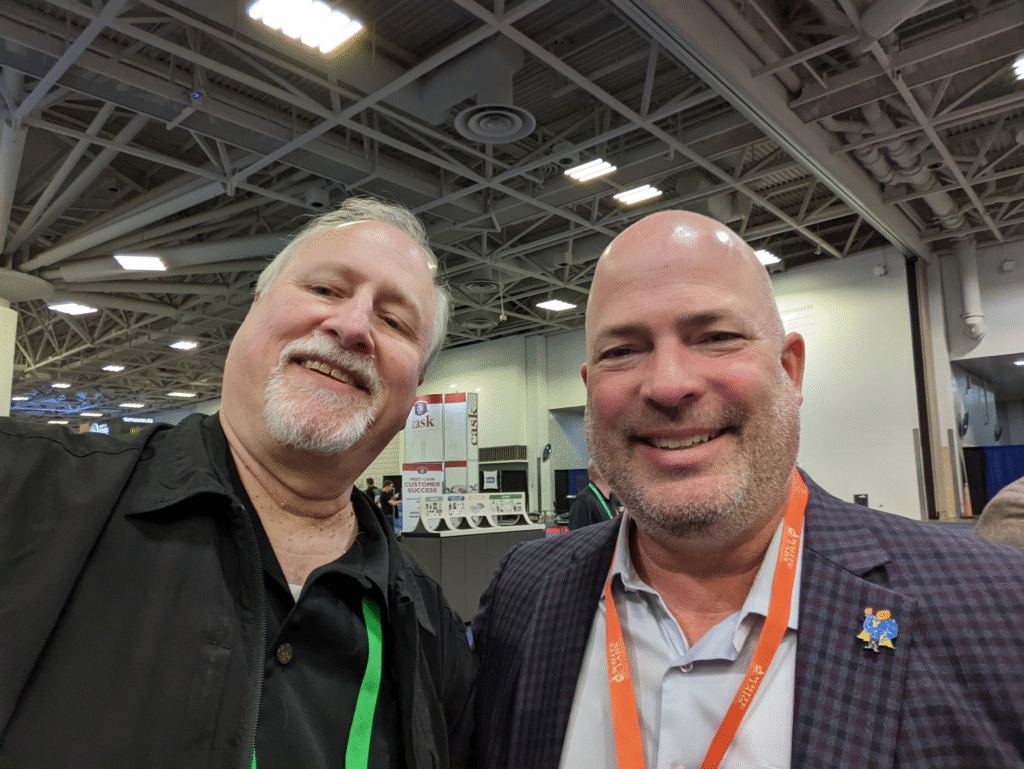





[Note: last two photos purloined from Facebook.]
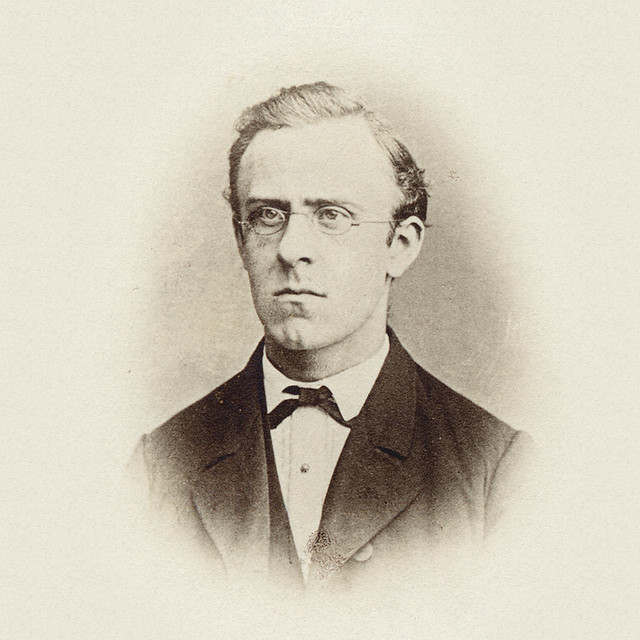
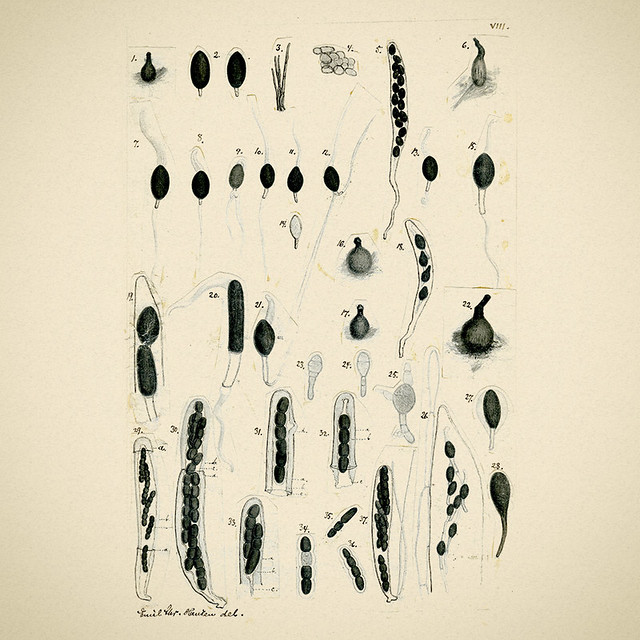






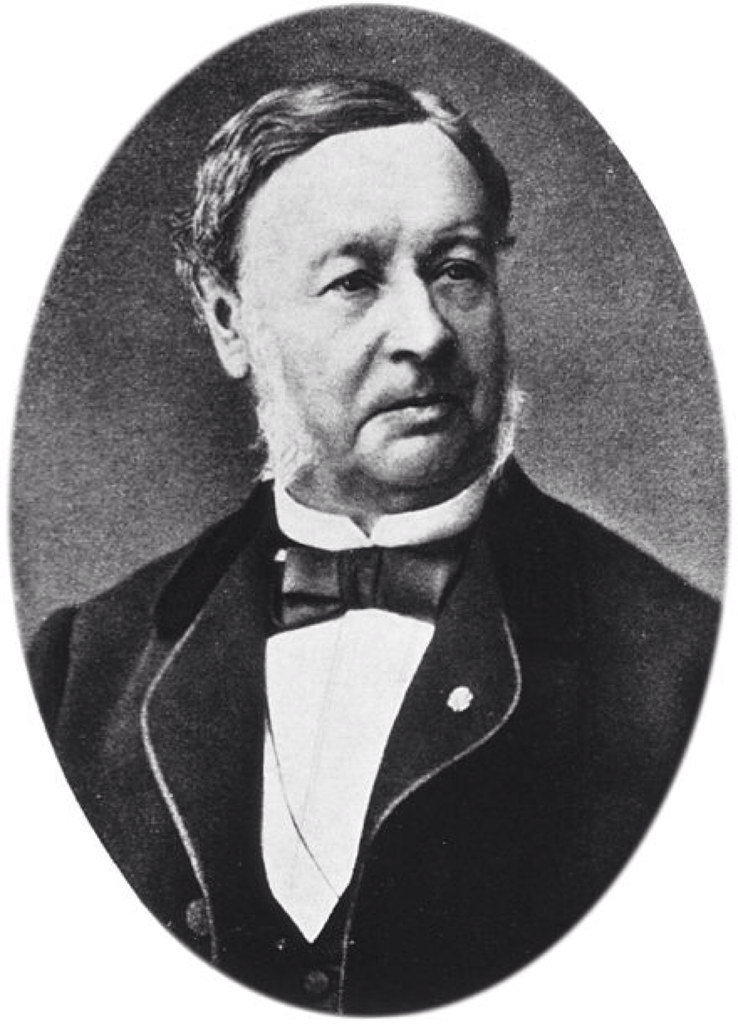
 Emil Christian Hansen, taken in 1908, a year before his death.
Emil Christian Hansen, taken in 1908, a year before his death.